Determination of Amino Acids in Feed by Pre-column Derivatization HPLC (AAP Method)
- Analyte: 17 Amino Acids (Asp, Glu, Ser, Gly, His, Arg, Thr, Ala, Pro, Tyr, Val, Met, Cys, Ile, Leu, Phe, Lys)
- System: EClassical 3100 HPLC System with UV Detector
- Column: EliteAAP Amino Acid Analysis Column (5 μm, 4.6 × 250 mm)
- Highlight: PITC pre-column derivatization HPLC method for simultaneous determination of 17 amino acids in feed with excellent linearity, reproducibility, and stability
Introduction
Amino acids are the fundamental units of proteins and essential nutrients for animal growth. Their composition and content are critical indicators of feed protein quality. However, due to the large number of amino acids, their similar structures, and the lack of strong UV absorption or fluorescence for most of them, amino acid analysis has always been challenging in HPLC. To address this, Elite (Dalian) Analytical Instruments Co., Ltd. has developed a complete solution for pre-column derivatization HPLC analysis of amino acids using phenyl isothiocyanate (PITC) as the derivatization reagent. The derivatization process is simple and rapid, yielding stable derivatives (up to 3 days) with high sensitivity, accuracy, and low cost, making it suitable for routine amino acid analysis in feed. The Elite-AAP system employs a dedicated C18 column to achieve baseline separation of 17 amino acids with UV detection at 254 nm.
Standards and Reagents
Amino acid standards: 17 amino acids (Asp, Glu, Ser, Gly, His, Arg, Thr, Ala, Pro, Tyr, Val, Met, Cys, Ile, Leu, Phe, Lys), purity ≥98%
Derivatization reagents: 1.0 mol/L triethylamine in acetonitrile, 0.2 mol/L phenyl isothiocyanate (PITC) in acetonitrile
Mobile phase components: mobile phase A: Acetonitrile/Methanol/Water=60:20:20 (v/v/v); mobile phase B: prepared from solid components (provided in the amino acid kit), dissolved in water, adjusted to pH 6.5±0.05 with glacial acetic acid, and diluted to volume. Filter through 0.45 μm membrane and degas before use.
Other reagents: Hydrochloric acid (6 mol/L, analytical grade), performic acid (freshly prepared, for oxidation of sulfur-containing amino acids if needed), n-Hexane (analytical grade), sodium acetate (for 0.05 mol/L sodium acetate solution), glacial acetic acid (for pH adjustment), acetonitrile, methanol (HPLC grade), deionized water (18.2 MΩ·cm)
Standard Solution Preparation
Amino acid stock solutions: Accurately weigh appropriate amounts of each amino acid standard and dissolve in 0.1 mol/L HCl to prepare individual stock solutions at concentrations of approximately 2.5 μmol/mL. Mix appropriate volumes to obtain a mixed amino acid stock solution.
Working standard solutions: Dilute the mixed stock solution with 0.1 mol/L HCl to obtain a series of concentrations covering the range 0.025–1.0 μmol/mL (e.g., 0.025, 0.25, and 1.0 μmol/mL). Take 200 μL of each working standard for derivatization following the same procedure as for samples.
Sample Pretreatment
- Oxidation (if determination of sulfur-containing amino acids is required): Weigh an appropriate amount of feed sample (equivalent to about 25 mg protein) into a 5 mL ampoule. Add 300 μL of freshly prepared, cooled performic acid, seal with parafilm, and place in an ice bath at 4°C for 16 h. Allow the sample to return to room temperature and evaporate to dryness under a gentle stream of nitrogen.
- Hydrolysis: Add 3 mL of 6 mol/L HCl to the ampoule, flame-seal the ampoule, and place in an oven at 110 ± 3°C for 24 h. After hydrolysis, cool to room temperature, open the ampoule, and evaporate the contents to dryness under nitrogen.
- Dissolution and dilution: Wash the ampoule repeatedly with small portions of water and transfer the combined washings to a 25 mL volumetric flask. Dilute to volume with water and mix well. Filter an aliquot through a 0.45 μm membrane filter.
- Derivatization: Pipette 200 μL of the filtered sample solution into a 1.5 mL plastic centrifuge tube. Add 100 μL of 1.0 mol/L triethylamine in acetonitrile and 100 μL of 0.2 mol/L PITC in acetonitrile. Vortex for 10 s and allow to stand at room temperature for 1 h.
Option A (for samples with high amino acid content): Add 0.6 mL of n-hexane, vortex for 1 min, and let stand for 10 min. Discard the upper hexane layer. Dilute the lower layer 4-fold with 0.05 mol/L sodium acetate solution.
Option B (for samples with low amino acid content): After derivatization, evaporate the mixture to dryness under nitrogen. Redissolve the residue in 400 μL (or as little as 100 μL for higher sensitivity) of 0.05 mol/L sodium acetate solution and vortex.
- Filter the derivatized solution through a 0.45 μm membrane filter before HPLC injection (10 μL)..
Instruments and Equipment
HPLC System: EClassical 3200 equipped with P3200 high-pressure pump, D3210 UV-Vis detector, S3210 autosampler, O3220 column oven, T3200 solvent bottle tray, Kromstation CDS.
Pretreatment equipment: Ultrasonic bath, solvent filtration apparatus, nitrogen evaporator, vortex mixer, pipettes (200 μL, 1 mL), drying oven (temperature >120°C), 5 mL ampoules, sealing film, 0.45 μm membrane filters (organic and aqueous), syringe filters, etc.
Chromatographic Conditions
- Column: EliteAAP amino acid analysis column (5 μm, 4.6 × 250 mm)
- Mobile Phase: A: Acetonitrile/Methanol/Water = 60 : 20 : 20 (v/v/v), degassed by sonication for 10–15 min
- B: Prepared from solid components (see "Mobile phase B" above), filtered through 0.45 μm membrane and degassed
- Flow Rate: 1.0 mL/min
- Detection: UV at 254 nm
- Injection Vol.: 20 μL
- Column Temp.: 42°C
Experimental Results
Standard Separation
A typical chromatogram of the 17 amino acid standards obtained under the described conditions is shown in Figure 1. All 17 amino acids are well resolved, meeting the requirements for qualitative and quantitative analysis.
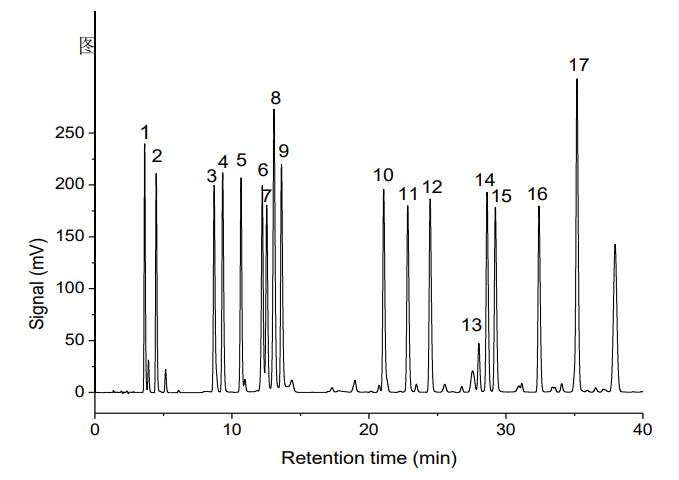
Fig. 1. Chromatogram of the 17 amino acid standards.
Peaks: 1-Asp, 2-Glu, 3-Ser, 4-Gly, 5-His, 6-Arg, 7-Thr, 8-Ala,
9-Pro, 10-Tyr, 11-Val, 12-Met, 13-Cys, 14-Ile, 15-Leu, 16-Phe, 17-Lys.
Method Performance
Linearity
Calibration curves were constructed using working standard solutions at concentrations of 0.025, 0.25, and 1.0 μmol/mL (and additional intermediate points). Linear regression equations and correlation coefficients for each amino acid are summarized in Table 1. All correlation coefficients (R) are greater than 0.9994, indicating excellent linearity over the tested range.
Table 1. Linear Equations and Correlation Coefficients for 17 Amino Acids
| Amino Acid |
Linear Equation (y = a x + b) |
Correlation Coefficient (R) |
| Asp |
y = 7,145.78 x – 103.85 |
0.9996 |
| Glu |
y = 7,210.30 x – 92.165 |
0.9997 |
| Ser |
y = 8,117.37 x – 64.25 |
0.9997 |
| Gly |
y = 8,205.19 x – 41.035 |
0.9996 |
| His |
y = 7,843.39 x – 107.435 |
0.9996 |
| Arg |
y = 5,985.01 x – 46.955 |
0.9997 |
| Thr |
y = 5,590.62 x – 42.57 |
0.9998 |
| Ala |
y = 12,971.29 x – 99.51 |
0.9997 |
| Pro |
y = 8,871.93 x + 9.20 |
0.9994 |
| Tyr |
y = 8,884.18 x – 102.57 |
0.9997 |
| Val |
y = 8,591.62 x – 67.24 |
0.9996 |
| Met |
y = 9,276.57 x – 85.38 |
0.9997 |
| Ile |
y = 8,946.21 x – 65.14 |
0.9996 |
| Leu |
y = 9,131.72 x – 90.24 |
0.9997 |
| Phe |
y = 8,837.57 x – 63.08 |
0.9997 |
| Lys |
y = 16,817.23 x – 94.68 |
0.9997 |
Detection limit
The lowest detectable concentration for aspartic acid (Asp) was found to be 3.13×10⁻⁶ mol/L. Similar detection limits are expected for other amino acids under the same conditions.
Repeatability
The method repeatability was evaluated by analyzing five replicate samples at three concentration levels (0.025, 0.25, and 1.0 μmol/mL). The relative standard deviations (RSD%) of peak areas for all 17 amino acids are summarized below: at 0.025 μmol/mL, RSD ranged from 1.96% to 4.90%; at 0.25 μmol/mL, RSD ranged from 1.00% to 1.99%; at 1.0 μmol/mL, RSD ranged from 0.87% to 1.76%. These results demonstrate excellent repeatability, especially at higher concentrations.
Accuracy
Accuracy was assessed by spiking experiments at a concentration of 0.25 μmol/mL (n=5). Recovery rates for the 17 amino acids ranged from 87.32% (Thr) to 118.48% (Asp). The complete recovery data are shown in Table 2.
| Amino Acid |
Recovery (%) |
Amino Acid |
Recovery (%) |
| Asp |
118.48 |
Pro |
109.48 |
| Glu |
106.68 |
Tyr |
90.44 |
| Ser |
94.17 |
Val |
93.93 |
| Gly |
104.06 |
Met |
106.77 |
| His |
96.97 |
Ile |
96.40 |
| Arg |
102.19 |
Leu |
100.80 |
| Thr |
87.32 |
Phe |
91.16 |
| Ala |
98.03 |
Lys |
95.74 |
Table 2. Linear equations and correlation coefficients for 17 amino acids
Derivative Stability
The stability of the PITC-derivatized amino acids was evaluated by analyzing the same sample over three consecutive days (three injections per day, n=9). The RSD% of peak areas for all amino acids was below 2% except for cysteine (Cys), which showed an RSD of 6.63%. The results confirm that the derivatives are stable for at least three days under the described conditions.

